Basic use
Ultra-short cheat sheet:
from datamatrix import DataMatrix, io
# Read a DataMatrix from file
dm = io.readtxt('data.csv')
# Create a new DataMatrix
dm = DataMatrix(length=5)
# The first two rows
print(dm[:2])
# Create a new column and initialize it with the Fibonacci series
dm.fibonacci = 0, 1, 1, 2, 3
# You can also specify column names as if they are dict keys
dm['fibonacci'] = 0, 1, 1, 2, 3
# Remove 0 and 3 with a simple selection
dm = (dm.fibonacci > 0) & (dm.fibonacci < 3)
# Get a list of indices that match certain criteria
print(dm[(dm.fibonacci > 0) & (dm.fibonacci < 3)])
# Select 1, 1, and 2 by matching any of the values in a set
dm = dm.fibonacci == {1, 2}
# Select all odd numbers with a lambda expression
dm = dm.fibonacci == (lambda x: x % 2)
# Change all 1s to -1
dm.fibonacci[dm.fibonacci == 1] = -1
# The first two cells from the fibonacci column
print(dm.fibonacci[:2])
# Column mean
print('Mean: %s' % dm.fibonacci.mean)
# Multiply all fibonacci cells by 2
dm.fibonacci_times_two = dm.fibonacci * 2
# Loop through all rows
for row in dm:
print(row.fibonacci) # get the fibonacci cell from the row
# Loop through all columns
for colname, col in dm.columns:
for cell in col: # Loop through all cells in the column
print(cell) # do something with the cell
# Or just see which columns exist
print(dm.column_names)
Important note: Because of a limitation (or feature, if you will) of the Python language, the behavior of and, or, and chained (x < y < z) comparisons cannot be modified. These therefore do not work with DataMatrix objects as you would expect them to:
# INCORRECT: The following does *not* work as expected
dm = dm.fibonacci > 0 and dm.fibonacci < 3
# INCORRECT: The following does *not* work as expected
dm = 0 < dm.fibonacci < 3
# CORRECT: Use the '&' operator
dm = (dm.fibonacci > 0) & (dm.fibonacci < 3)
Slightly longer cheat sheet:
- Basic operations
- Creating a DataMatrix
- Concatenating two DataMatrix objects
- Creating columns
- Renaming columns
- Deleting columns
- Slicing and assigning to column cells
- Column properties
- Iterating over rows, columns, and cells
- Selecting data
- Comparing a column to a value
- Selecting by multiple criteria with | (or), & (and), and ^ (xor)
- Selecting by multiple criteria by comparing to a set {}
- Selecting with a function or lambda expression
- Selecting values that match another column (or sequence)
- Getting indices for rows that match selection criteria ('where')
- Element-wise column operations
- Column types
- Reading and writing files
Basic operations
Creating a DataMatrix
Create a new DataMatrix object, and add a column (named col). By default, the column is of the MixedColumn type, which can store numeric and string data.
import sys
from datamatrix import DataMatrix, __version__
dm = DataMatrix(length=2)
dm.col = ':-)'
print(
'Examples generated with DataMatrix v{} on Python {}\n'.format(
__version__,
sys.version
)
)
print(dm)
Output:
Examples generated with DataMatrix v0.15.0 on Python 3.10.5 | packaged by conda-forge | (main, Jun 14 2022, 07:04:59) [GCC 10.3.0]
+---+-----+
| # | col |
+---+-----+
| 0 | :-) |
| 1 | :-) |
+---+-----+
You can change the length of the DataMatrix later on. If you reduce the length, data will be lost. If you increase the length, empty cells will be added.
dm.length = 3
Concatenating two DataMatrix objects
You can concatenate two DataMatrix objects using the << operator. Matching columns will be combined. (Note that row 2 is empty. This is because we have increased the length of dm in the previous step, causing an empty row to be added.)
dm2 = DataMatrix(length=2)
dm2.col = ';-)'
dm2.col2 = 10, 20
dm3 = dm << dm2
print(dm3)
Output:
+---+-----+------+
| # | col | col2 |
+---+-----+------+
| 0 | :-) | |
| 1 | :-) | |
| 2 | | |
| 3 | ;-) | 10 |
| 4 | ;-) | 20 |
+---+-----+------+
Creating columns
You can change all cells in column to a single value. This creates a new column if it doesn't exist yet.
dm.col = 'Another value'
print(dm)
Output:
+---+---------------+
| # | col |
+---+---------------+
| 0 | Another value |
| 1 | Another value |
| 2 | Another value |
+---+---------------+
You can change all cells in a column based on a sequence. This creates a new column if it doesn't exist yet. This sequence must have the same length as the column (3 in this case).
dm.col = 1, 2, 3
print(dm)
Output:
+---+-----+
| # | col |
+---+-----+
| 0 | 1 |
| 1 | 2 |
| 2 | 3 |
+---+-----+
If you do not know the name of a column, for example because it is defined by a variable, you can also refer to columns as though they are items of a dict. However, this is not recommended, because it makes it less clear whether you are referring to column or a row.
dm['col'] = 'X'
print(dm)
Output:
+---+-----+
| # | col |
+---+-----+
| 0 | X |
| 1 | X |
| 2 | X |
+---+-----+
Renaming columns
dm.rename('col', 'col2')
print(dm)
Output:
+---+------+
| # | col2 |
+---+------+
| 0 | X |
| 1 | X |
| 2 | X |
+---+------+
Deleting columns
You can delete a column using the del keyword:
dm.col = 'x'
del dm.col2
print(dm)
Output:
+---+-----+
| # | col |
+---+-----+
| 0 | x |
| 1 | x |
| 2 | x |
+---+-----+
Slicing and assigning to column cells
Assign to one cell
dm.col[1] = ':-)'
print(dm)
Output:
+---+-----+
| # | col |
+---+-----+
| 0 | x |
| 1 | :-) |
| 2 | x |
+---+-----+
Assign to multiple cells
This changes row 0 and 2. It is not a slice!
dm.col[0,2] = ':P'
print(dm)
Output:
+---+-----+
| # | col |
+---+-----+
| 0 | :P |
| 1 | :-) |
| 2 | :P |
+---+-----+
Assign to a slice of cells
dm.col[1:] = ':D'
print(dm)
Output:
+---+-----+
| # | col |
+---+-----+
| 0 | :P |
| 1 | :D |
| 2 | :D |
+---+-----+
Assign to cells that match a selection criterion
dm.col[1:] = ':D'
dm.is_happy = 'no'
dm.is_happy[dm.col == ':D'] = 'yes'
print(dm)
Output:
+---+-----+----------+
| # | col | is_happy |
+---+-----+----------+
| 0 | :P | no |
| 1 | :D | yes |
| 2 | :D | yes |
+---+-----+----------+
Column properties
Basic numeric properties, such as the mean, can be accessed directly. Only numeric values are taken into account.
dm.col = 1, 2, 'not a number'
# Numeric descriptives
print('mean: %s' % dm.col.mean)
print('median: %s' % dm.col.median)
print('standard deviation: %s' % dm.col.std)
print('sum: %s' % dm.col.sum)
print('min: %s' % dm.col.min)
print('max: %s' % dm.col.max)
# Other properties
print('unique values: %s' % dm.col.unique)
print('number of unique values: %s' % dm.col.count)
print('column name: %s' % dm.col.name)
Output:
mean: 1.5
median: 1.5
standard deviation: 0.7071067811865476
sum: 3.0
min: 1.0
max: 2.0
unique values: [1, 2, 'not a number']
number of unique values: 3
column name: col
Iterating over rows, columns, and cells
By iterating directly over a DataMatrix object, you get successive Row objects. From a Row object, you can directly access cells.
dm.col = 'a', 'b', 'c'
for row in dm:
print(row)
print(row.col)
Output:
+----------+-------+
| Name | Value |
+----------+-------+
| col | a |
| is_happy | no |
+----------+-------+
a
+----------+-------+
| Name | Value |
+----------+-------+
| col | b |
| is_happy | yes |
+----------+-------+
b
+----------+-------+
| Name | Value |
+----------+-------+
| col | c |
| is_happy | yes |
+----------+-------+
c
By iterating over DataMatrix.columns, you get successive (column_name, column) tuples.
for colname, col in dm.columns:
print('%s = %s' % (colname, col))
Output:
col = col['a', 'b', 'c']
is_happy = col['no', 'yes', 'yes']
By iterating over a column, you get successive cells:
for cell in dm.col:
print(cell)
Output:
a
b
c
By iterating over a Row object, you get (column_name, cell) tuples:
row = dm[0] # Get the first row
for colname, cell in row:
print('%s = %s' % (colname, cell))
Output:
col = a
is_happy = no
The column_names property gives a sorted list of all column names (without the corresponding column objects):
print(dm.column_names)
Output:
['col', 'is_happy']
Selecting data
Comparing a column to a value
You can select by directly comparing columns to values. This returns a new DataMatrix object with only the selected rows.
dm = DataMatrix(length=10)
dm.col = range(10)
dm_subset = dm.col > 5
print(dm_subset)
Output:
+---+-----+
| # | col |
+---+-----+
| 6 | 6 |
| 7 | 7 |
| 8 | 8 |
| 9 | 9 |
+---+-----+
Selecting by multiple criteria with | (or), & (and), and ^ (xor)
You can select by multiple criteria using the | (or), & (and), and ^ (xor) operators (but not the actual words 'and' and 'or'). Note the parentheses, which are necessary because |, &, and ^ have priority over other operators.
dm_subset = (dm.col < 1) | (dm.col > 8)
print(dm_subset)
Output:
+---+-----+
| # | col |
+---+-----+
| 0 | 0 |
| 9 | 9 |
+---+-----+
dm_subset = (dm.col > 1) & (dm.col < 8)
print(dm_subset)
Output:
+---+-----+
| # | col |
+---+-----+
| 2 | 2 |
| 3 | 3 |
| 4 | 4 |
| 5 | 5 |
| 6 | 6 |
| 7 | 7 |
+---+-----+
Selecting by multiple criteria by comparing to a set {}
If you want to check whether column values are identical to, or different from, a set of test values, you can compare the column to a set object. (This is considerably faster than comparing the column values to each of the test values separately, and then merging the result using & or |.)
dm_subset = dm.col == {1, 3, 5, 7}
print(dm_subset)
Output:
+---+-----+
| # | col |
+---+-----+
| 1 | 1 |
| 3 | 3 |
| 5 | 5 |
| 7 | 7 |
+---+-----+
Selecting with a function or lambda expression
You can also use a function or lambda expression to select column values. The function must take a single argument and its return value determines whether the column value is selected. This is analogous to the classic filter() function.
dm_subset = dm.col == (lambda x: x % 2)
print(dm_subset)
Output:
+---+-----+
| # | col |
+---+-----+
| 1 | 1 |
| 3 | 3 |
| 5 | 5 |
| 7 | 7 |
| 9 | 9 |
+---+-----+
Selecting values that match another column (or sequence)
You can also select by comparing a column to a sequence, in which case a row-by-row comparison is done. This requires that the sequence has the same length as the column, is not a set object (because set objects are treated as described above).
dm = DataMatrix(length=4)
dm.col = 'a', 'b', 'c', 'd'
dm_subset = dm.col == ['a', 'b', 'x', 'y']
print(dm_subset)
Output:
+---+-----+
| # | col |
+---+-----+
| 0 | a |
| 1 | b |
+---+-----+
When a column contains values of different types, you can also select values by type: (Note: On Python 2, all str values are automatically decoded to unicode, so you'd need to compare the column to unicode to extract str values.)
dm = DataMatrix(length=4)
dm.col = 'a', 1, 'c', 2
dm_subset = dm.col == int
print(dm_subset)
Output:
+---+-----+
| # | col |
+---+-----+
| 1 | 1 |
| 3 | 2 |
+---+-----+
Getting indices for rows that match selection criteria ('where')
You can get the indices for rows that match certain selection criteria by slicing a DataMatrix with a subset of itself. This is similar to the numpy.where() function.
dm = DataMatrix(length=4)
dm.col = 1, 2, 3, 4
print(dm[(dm.col > 1) & (dm.col < 4)])
Output:
[1, 2]
Element-wise column operations
Multiplication, addition, etc.
You can apply basic mathematical operations on all cells in a column simultaneously. Cells with non-numeric values are ignored, except by the + operator, which then results in concatenation.
dm = DataMatrix(length=3)
dm.col = 0, 'a', 20
dm.col2 = dm.col*.5
dm.col3 = dm.col+10
dm.col4 = dm.col-10
dm.col5 = dm.col/50
print(dm)
Output:
+---+-----+------+------+------+------+
| # | col | col2 | col3 | col4 | col5 |
+---+-----+------+------+------+------+
| 0 | 0 | 0.0 | 10 | -10 | 0.0 |
| 1 | a | a | a10 | a | a |
| 2 | 20 | 10.0 | 30 | 10 | 0.4 |
+---+-----+------+------+------+------+
Applying a function or lambda expression
@ operator is only available in Python 3.5 and later.
You can apply a function or lambda expression to all cells in a column simultaneously with the @ operator.
dm = DataMatrix(length=3)
dm.col = 0, 1, 2
dm.col2 = dm.col @ (lambda x: x*2)
print(dm)
Output:
+---+-----+------+
| # | col | col2 |
+---+-----+------+
| 0 | 0 | 0 |
| 1 | 1 | 2 |
| 2 | 2 | 4 |
+---+-----+------+
Column types
When you create a DataMatrix, you can indicate a default column type. If you do not specify a default column type, a MixedColumn is used by default.
from datamatrix import DataMatrix, IntColumn
dm = DataMatrix(length=2, default_col_type=IntColumn)
dm.i = 1, 2 # This is an IntColumn
You can also explicitly indicate the column type when creating a new column:
from datamatrix import FloatColumn
dm.f = FloatColumn
MixedColumn (default)
A MixedColumn contains text (unicode in Python 2, str in Python 3), int, float, or None.
Important notes:
utf-8encoding is assumed for byte strings- String with numeric values, including
NANandINF, are automatically converted to the most appropriate type - The string 'None' is not converted to the type
None - Trying to assign a non-supported type results in a
TypeError
from datamatrix import DataMatrix, NAN, INF
dm = DataMatrix(length=12)
dm.datatype = (
'int',
'int (converted)',
'float',
'float (converted)',
'None',
'str',
'float',
'float (converted)',
'float',
'float (converted)',
'float',
'float (converted)',
)
dm.value = (
1,
'1',
1.2,
'1.2',
None,
'None',
NAN,
'nan',
INF,
'inf',
-INF,
'-inf'
)
print(dm)
Output:
+----+-------------------+-------+
| # | datatype | value |
+----+-------------------+-------+
| 0 | int | 1 |
| 1 | int (converted) | 1 |
| 2 | float | 1.2 |
| 3 | float (converted) | 1.2 |
| 4 | None | None |
| 5 | str | None |
| 6 | float | nan |
| 7 | float (converted) | nan |
| 8 | float | INF |
| 9 | float (converted) | INF |
| 10 | float | -inf |
| 11 | float (converted) | -inf |
+----+-------------------+-------+
IntColumn (requires numpy)
The IntColumn contains only int values. As of 0.14, the easiest way to create a IntColumn column is to assign int to a new column name.
Important notes:
- Trying to assign a value that cannot be converted to an
intresults in aTypeError - Float values will be rounded down (i.e. the decimals will be lost)
NANorINFvalues are not supported because these arefloat
from datamatrix import DataMatrix
dm = DataMatrix(length=2)
dm.i = int
dm.i = 1, 2
print(dm)
Output:
+---+---+
| # | i |
+---+---+
| 0 | 1 |
| 1 | 2 |
+---+---+
If you insert non-int values, they are automatically converted to int if possible. Decimals are discarded (i.e. values are floored, not rounded):
dm.i = '3', 4.7
print(dm)
Output:
+---+---+
| # | i |
+---+---+
| 0 | 3 |
| 1 | 4 |
+---+---+
If you insert values that cannot converted to int, a TypeError is raised:
try:
dm.i = 'x'
except TypeError as e:
print(repr(e))
Output:
TypeError('IntColumn expects integers, not x')
FloatColumn (requires numpy)
The FloatColumn contains float, nan, and inf values. As of 0.14, the easiest way to create a FloatColumn column is to assign float to a new column name.
Important notes:
- Values that are accepted by a
MixedColumnbut cannot be converted to a numeric value becomeNAN. Examples are non-numeric strings orNone. - Trying to assign a non-supported type results in a
TypeError
import numpy as np
from datamatrix import DataMatrix, FloatColumn
dm = DataMatrix(length=3)
dm.f = float
dm.f = 1, np.nan, np.inf
print(dm)
Output:
+---+-----+
| # | f |
+---+-----+
| 0 | 1.0 |
| 1 | nan |
| 2 | INF |
+---+-----+
If you insert other values, they are automatically converted if possible.
dm.f = '3.3', 'inf', 'nan'
print(dm)
Output:
+---+-----+
| # | f |
+---+-----+
| 0 | 3.3 |
| 1 | INF |
| 2 | nan |
+---+-----+
If you insert values that cannot be converted to float, they become nan.
dm.f = 'x'
print(dm)
Output:
/home/sebastiaan/anaconda3/envs/pydata/lib/python3.10/site-packages/datamatrix/py3compat.py:105: UserWarning: Invalid type for
FloatColumn: x
warnings.warn(safe_str(msg), *args)
[32m⠙[0m Generating...+---+-----+
| # | f |
+---+-----+
| 0 | nan |
| 1 | nan |
| 2 | nan |
+---+-----+
nan data!
You have to take special care when working with nan data. In general, nan is not equal to anything else, not even to itself: nan != nan. You can see this behavior when selecting data from a FloatColumn with nan values in it.
from datamatrix import DataMatrix, FloatColumn
dm = DataMatrix(length=3)
dm.f = FloatColumn
dm.f = 0, np.nan, 1
dm = dm.f == [0, np.nan, 1]
print(dm)
Output:
+---+-----+
| # | f |
+---+-----+
| 0 | 0.0 |
| 2 | 1.0 |
+---+-----+
However, for convenience, you can select all nan values by comparing a FloatColumn to a single nan value:
from datamatrix import DataMatrix, FloatColumn
dm = DataMatrix(length=3)
dm.f = FloatColumn
dm.f = 0, np.nan, 1
print('NaN values')
print(dm.f == np.nan)
print('Non-NaN values')
print(dm.f != np.nan)
Output:
NaN values
+---+-----+
| # | f |
+---+-----+
| 1 | nan |
+---+-----+
Non-NaN values
+---+-----+
| # | f |
+---+-----+
| 0 | 0.0 |
| 2 | 1.0 |
+---+-----+
SeriesColumn: Working with continuous data (requires numpy)
The SeriesColumn is 2 dimensional; that is, each cell is by itself an array of values. Therefore, the SeriesColumn can be used to work with sets of continuous data, such as EEG or eye-position traces.
For more information about series, see:
import numpy as np
from matplotlib import pyplot as plt
from datamatrix import SeriesColumn
length = 10 # Number of traces
depth = 50 # Size of each trace
x = np.linspace(0, 2*np.pi, depth)
sinewave = np.sin(x)
noise = np.random.random(depth)*2-1
dm = DataMatrix(length=length)
dm.series = SeriesColumn(depth=depth)
dm.series[0] = noise
dm.series[1:].setallrows(sinewave)
dm.series[1:] *= np.linspace(-1, 1, 9)
plt.xlim(x.min(), x.max())
plt.plot(x, dm.series.plottable, color='green', linestyle=':')
y1 = dm.series.mean-dm.series.std
y2 = dm.series.mean+dm.series.std
plt.fill_between(x, y1, y2, alpha=.2, color='blue')
plt.plot(x, dm.series.mean, color='blue')
plt.show()
Output:
/home/sebastiaan/anaconda3/envs/pydata/lib/python3.10/site-packages/numpy/lib/nanfunctions.py:1879: RuntimeWarning: Degrees of
freedom <= 0 for slice.
var = nanvar(a, axis=axis, dtype=dtype, out=out, ddof=ddof,
[32m⠴[0m Generating...
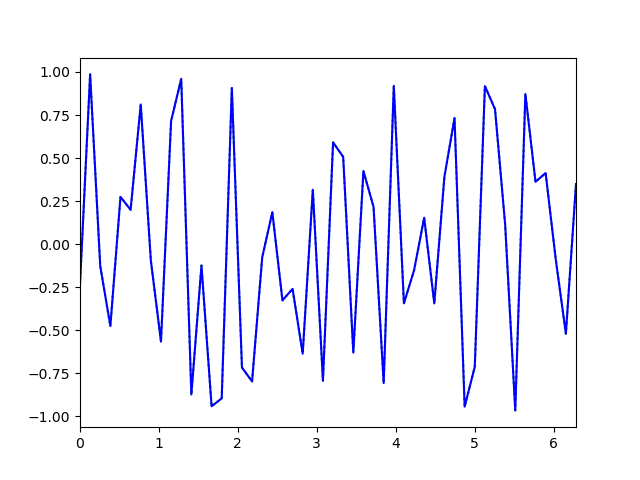
You can also create a SeriesColumn by assigning a 2D numpy array to a new column, where one of the dimensions matches the length of the DataMatrix. The other dimension is then assumed to be the depth of the SeriesColumn:
dm = DataMatrix(length=3)
dm.random_noise = np.random.random((3, 10))
Reading and writing files
You can read and write files with functions from the datamatrix.io module. The main supported file types are csv and xlsx.
from datamatrix import io
dm = DataMatrix(length=3)
dm.col = 1, 2, 3
# Write to disk
io.writetxt(dm, 'my_datamatrix.csv')
io.writexlsx(dm, 'my_datamatrix.xlsx')
# And read it back from disk!
dm = io.readtxt('my_datamatrix.csv')
dm = io.readxlsx('my_datamatrix.xlsx')



